Documentation
Table of Contents
API
Right now, the API is very minimal, the current access is just enough to give the basic functionality to the entire site. These features include: dynamic generation of .gjf, .mol2, .svg, and .png files for any molecule given the name.
Naming
For generating the molecules, there is a very rough Finite State Machine that parses through the names given and spits out either the molecule requested or an error. Here is roughly what the context free grammar would look like.
digit = "0" | "1" | "2" | "3" | "4" | "5" | "6" | "7" | "8" | "9" ;
int = digit, { digit } ;
YY = "N" | "P" | "C" ;
XX = "O" | "S" | YY ;
type = "C" | "T" | "E" | "Z" ;
core = type, XX, YY ;
aryl0 = "10" | "11" | "2" | "3" | "8" | "9" ;
aryl2 = "12" | "13" | "4" | "5" | "6" | "7" ;
xgroup = "A" | "B" | "C" | "D" | "E" | "F" | "G" | "H" | "I" | "J" | "K" | "L" | "M" ;
rgroup = "a" | "b" | "c" | "d" | "e" | "f" | "g" | "h" | "i" | "j" | "k" | "l" | "m" ;
rotate = "-" | "(", int, ")"
arylfill = aryl2, [rotate], [rgroup], [rotate], [rgroup], [rotate] ;
allaryl = aryl0 | arylfill ;
end = [allaryl, {allaryl}], [xgroup] ;
extend = ("n" | "m"), int ;
stack = ("x" | "y" | "z"), int ;
meta = ["_", extend], ["_", stack] {"_", stack} ;
chain = end, {end}, [meta] ;
rbenzo = [end, "_"], core, ["_", end, ["_"]], [("_" | "__"), end] ;
lbenzo =
multibenzo = [end, "_"], core, ["_", end, ["_"]], [("_" | "__"), end] ;
molecule = chain | benzo | multibenzo ;
Molecule Specific
For all of the molecules there is a basic API access to allow getting the different types of outputs. The first, and most common, being the gjf output. This is the standard Gaussian file type and is what should be used for running the calculations. There is also an added possible parameter called "keywords" that can be added to add/change the keywords/settings of the molecule. If none is given, then opt B3LYP/6-31g(d) is assumed. Another possible parameter is "view". If this is enabled the output will be browser viewable rather than a download.
/chem/$NAME.gjf
/chem/$NAME.gjf?keywords=B3LYP/6-31g(d)
/chem/$NAME.gjf?view=true
The next form of output is the mol2 format. This is added because it is a fairly simple interchange format for different software packages.
/chem/$NAME.mol2
/chem/$NAME.mol2?view=true
The last type of molecule specific access is an image of the structure. This comes in two different forms, both of which show the exact same thing. The first is a standard .png image. The second is a vector version in the form of a .svg. It is a very basic rendering over the overall structure of the molecule.
Single Bond = single black line
Aromatic Bond = two dashed red lines
Double Bond = two green lines
Triple Bond = three blue lines
Sulfur = yellow dot
Oxygen = red dot
Nitrogen = blue dot
Chlorine = green dot
Carbon = medium gray dot
Phosphorus = orange dot
Hydrogen = off white dot
Silicon = green/gray dot
Flourine = green/blue dot
Similar to the gjf file, the images can be parameterized, with their scaling. The default view is a size 10 which means the atoms have a diameter of 10.
/chem/$NAME.png
/chem/$NAME.png?size=20
/chem/$NAME.svg
/chem/$NAME.svg?size=20
The whole thing is very hackish and is just intended to allow a preview of the molecule without having to open it in Gaussian. As expected of a 2D Image, three dimensionality is poorly shown. this is especially apparent in molecules with TMS or Carbazole. This is also compounded with the fact that the fragments have a hackish transform to align them)
/static/docs/img/7k_TON_7k_7k.png
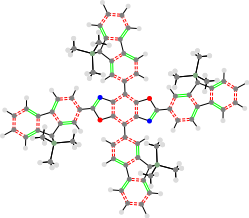
Molecule JSON Data
If you want a simple machine readable collection of the properties of a given name you can use the molecule JSON interface. Many of the values that are returned by this are dependent on the name of the molecule and if the calculated values of the molecule are already in the database. property_predictions will only be available for names that fit the subset of the naming scheme that the machine learning was done (single core). The property_limits values will be available if the name fits the subset of the naming scheme and if that respective direction can be polymerized (ie, no X-Groups capping expansion in that direction). datapoint will only be seen in structures that already have calculated values stored in the database.
{
"features": [
[ <NAIVE FEATURE VECTOR> ],
[ <DECAY FEATURE VECTOR> ],
],
"molecule": "4a_TON",
"property_predictions": [-5.8055209124754841, -1.9840131601722049, 3.649533471581226],
"exact_name": "4aaA_TON_A_A_n1_m1_x1_y1_z1",
"exact_name_spacers": "4aaA**_TON_A**_A**_n1_m1_x1_y1_z1",
"property_limits": {
"m": [-5.6012581484079291, -2.2255173089165798, 3.0106583051116274],
"n": [-5.8039781252330664, -2.8912498957468986, 2.7177841447390274]
},
"structure_type": "Benzobisazole",
"name_error": null,
"error_report": null,
"new": false,
"datapoint": {
"band_gap": 4.0578,
"name": "4_TON_A_A",
"energy": -798.2647132,
"homo_orbital": 61,
"id": 1882,
"lumo": -1.68737791307,
"homo": -5.99983802696,
"exact_name": "4aaA_TON_A_A_n1_m1_x1_y1_z1",
"dipole": 1.3283,
"options": "td B3LYP/6-31g(d)"},
"exact_name_spacers": "4aaA**_TON_A**_A**_n1_m1_x1_y1_z1"
}
}
In addition to the standard information about the molecules, one can also get the geometry in a JSON format as seen below.
{
"bonds": [
{
"second": 1,
"type": "1",
"first": 0
},
{
"second": 2,
"type": "2",
"first": 0
},
{
"second": 5,
"type": "1",
"first": 0
},
{
"second": 3,
"type": "1",
"first": 2
},
{
"second": 4,
"type": "1",
"first": 2
}
],
"center": [
-0.33813333333333,
0.58133333333333,
0.00015
],
"atoms": [
{
"y": 0,
"x": 0,
"z": 0,
"element": "C"
},
{
"y": 0,
"x": 1.0867,
"z": 0,
"element": "H"
},
{
"y": 1.1717,
"x": -0.6771,
"z": 0,
"element": "C"
},
{
"y": 1.1457,
"x": -1.7636,
"z": 0.0015,
"element": "H"
},
{
"y": 2.3642,
"x": -0.1596,
"z": -0.0006,
"element": "H"
},
{
"y": -1.1936,
"x": -0.5152,
"z": 0,
"element": "H"
}
]
}
Jobs
Jobs have a few required parameters: name, email, cluster, nodes, and walltime. name and email are just as they seem. The former being the name of the job/file (this can be used to setup time dependent files). The latter just being the email to send the job updates to. cluster corresponds to the single letter at the start of the cluster's name.
Gordon = g
Blacklight = b
Trestles = t
Hooper = h
Carver = c
Localhost = l
nodes is the number of nodes to use on the cluster. This value is multiplied by 16 for the clusters that require ncpu numbers instead of nodes. The final value walltime is the maximum amount of time, in hours, that the job should run.
When submitting jobs, it returns a bit of information to tell the state of the jobs submission. This comes in the form of a simple json object with two values. error and jobid. If error is not null then that means the job submission failed and it will display the error that occurred. Otherwise jobid will display the number of the job submitted.
{
"jobid": 123,
"error": null,
}
Multi Molecule
The multi molecule view works much as you might expect with molecule names comma delimited. This is useful when looking at just a couple of molecules.
/chem/$NAME1,$NAME2,$NAME3/
This method makes it simple to make a few nonrelated molecules quickly.
/chem/24a_TON,35b_TNN,4g_CCC_4g/
Just like with single molecules, it is possible to set the keywords to be something other than B3LYP/6-31g(d).
/chem/$NAME1,$NAME2,$NAME3/?keywords=HF/3-21G
In addition to getting all of the structures given, there is also the ability to randomly sample the structures using the random parameter. The following will return two random structures from the three given.
/chem/$NAME1,$NAME2,$NAME3/?random=2
Brace Expansion
Along with comma separated names, there is also an added feature that works much like Unix brace expansion.
Example:
$ echo test{ing,er,ed,}
testing tester tested test
$ echo {a,b,c}{a,b,c}
aa ab ac ba bb bc ca cb cc
In the case of chemtools-webapp, the usage is much the same.
Variables
Along with that functionality, there are some added variables that can be accessed the same as shell variables.
Example:
$ echo $USER
chris
$ echo $HOME
/home/chris
With chemtools-webapp there are six variables each of which correspond to a set of the naming scheme.
CORES = "{C,T,E,Z}{O,S,N,P,C}{N,P,C}"
SCORES = "{E,Z}{O,S,N,P,C}{N,P,C}"
DCORES = "{C,T}{O,S,N,P,C}{N,P,C}"
RGROUPS = "a,b,c,d,e,f,g,h,i,j,k,l"
XGROUPS = "A,B,C,D,E,F,G,H,I,J,K,L"
ARYL = "2,3,4,5,6,7,8,9,10,11,12,13"
ARYL0 = "2,3,8,9,10,11"
ARYL2 = "4,5,6,7,12,13"
So, if you wanted to create all of the substituant combinations for 4X_TON, rather than typing all the substituants out, you can just use:
Or if you wanted all the R-groups and all the cores:
Internal Variables
Now, that may seem well and good, except in the case where you may have multiple parts that you want the same. Like with 4X_TON_4X_4X. In that case, there are some special variables that correspond to the number of the replacement.
/chem/4{$RGROUPS}_TON_4{$0}_4{$0}/
/chem/{$ARYL2}{$RGROUPS}_TON_{$0}{$1}_{$0}{$1}/
Currently, there is no way to simplify the name with heavy repetitions in it. An example being something like the first one below without major changes in the grammar. That being said, this method does make it trivial to make several thousand molecules in the matter of a second or two.
4X4X4X4X_TON_4X4X4X4X_4X4X4X4X
/chem/{$ARYL2}{$ARYL2}{$RGROUPS}_{$CORES}_{$0}{$1}{$2}_{$0}{$1}{$2}_n{1,2,3,4}/
This case creating 4 * 4 * 12 * 8 * 4 = 6144 molecules. Due to optimizations, generating the page with all of these molecules is trivial (within a second or so), generating the zip file with all of them in it; however, is not (~5 minutes and ~100 megs of just gjf files). Because of this, there is a arbitrary timeout limit when generating molecules of 1 second. This could be optimized later to at least seem more responsive, but it is not a concern because no one is going to be dealing with more than about 100 molecules at a time. With a reasonable case as follows. Which is completed in a fraction of a second for both the generation and the download.
/chem/4{$RGROUPS}_TON_4{$0}_4{$0}_n{1,2,3,4},4a_{$CORES}_4a_4a_n{1,2,3,4}/
In addition to the values of the there are also 2 options that can be applied to the variables to change the case. These are added by adding a .U (for uppercase) or a .L (for lowercase) to the end of the variable number. This is useful when making matching R-Groups and X-Groups.
/chem/4{$RGROUPS}_TON_4{$0}_{$0.U}/
Zip Output
Added with this main display page is another API type access to allow generating zip files with all the molecules of a given pattern/set. By default this will include all of the .gjf files for the molecules. The following two examples are equivalent.
/chem/$PATTERN.zip
/chem/$PATTERN.zip?gjf=true
In addition to being able to get gjf files this can also be used to download jobs, .mol2, and .png files of the molecules. This can be done by setting job, mol2, and image to true, respectively.
/chem/$PATTERN.zip?mol2=true
/chem/$PATTERN.zip?gjf=true
/chem/$PATTERN.zip?image=true&gjf=true
Note that the job option requires the inclusion of all of the job parameters.
/chem/$PATTERN.zip?job=true&name={{name}}&email=e@t.com&cluster=b&nodes=1&walltime=1
Any errors in the output will be written to a file called errors.txt.
Name Check
If you want to check the validity of a name or set of names you can use the name checker. The API call returns a json object with two values: molecules and error. error is used to keep track of problems when doing the entire set. Most time this is just a timeout error due to the query taking longer than an arbitrary limit of 1 second. The second value is a array of arrays. With each array corresponding to a molecule name. Within each one they come in the form [name, warning, error]. Errors are actual problems in the name, whereas warnings are user submitted problems.
{
"molecules": [
["24a_TON", true, null],
["24ball_TON", null, "no rgroups allowed at start"]
],
"error": null
}
Jobs
With multiple molecules this adds a small layer of complexity with respect to naming. This comes in the form of a generic naming variable {{ name }}. So for example, if you wanted to create all the time dependent job names. The following two names would be equivalent.
{{ name }}_TD
{{name}}_TD
Just like with single molecules, at this time there is Alpha functionality to be able submit jobs to the clusters. When the jobs are submitted a bit of json will be returned to display the status of the jobs. There are two main lists, the first being failed. It corresponds to the jobs that failed for some reason. Each item in the list is a list of the name of the job and the reason why it failed. The second list is a list of name and jobid pairs. The last value returned is an overall error value that is only non-null if the person trying to submit is not a staff user.
POST /chem/24{a,ball}_TON?name={{name}}&email=e@t.com&cluster=b&nodes=1&walltime=1
{
"failed": [
["24ball_TON", "no rgroups allowed"],
...
...
],
"worked" : [
["24a_TON", 1000],
...
...
],
"error" : null,
}
Fragments
All of the fragments used in generating the molecules can be found here:
/chem/frag/$NAME/
They use a slightly altered XYZ file format in the form of:
Element0 x0 y0 z0
Element1 x1 y1 z1
...
ElementN xN yN zN
Atom0 Atom1 Connection0
Atom1 Atom2 Connection1
...
AtomN-1 AtomN ConnectionN
Where x, y, and z are all floats. Element is a String from the set of all the elements plus the addition of a few special characters to signify where to bond to. Atom1 and Atom2 are both Integers corresponding to the location of the atom in the coordinate list. The connection is a string in the set ["1", "2", "3", "Ar"], where 1, 2, and 3 are single, double and triple bonds, respectively. Ar is an Aromatic (1.5 bond).
Here is an example of the Triple Bond.
C 0.402800 -0.479100 -0.000100
C 1.209100 0.607700 -0.000300
~0 2.383000 2.189800 -0.000700
~1 -0.747300 -2.029000 0.000300
1 2 3
1 4 Ar
2 3 Ar
There are 3 added symbols in the character set for the element names and those are "~", "*", and "+". These are used to signify the type of things the element can bond to. After the set of possible things to bond to is a number that indicates the order that the bonds get used. So, in the case of the cores, the correct parts of the molecule are built in the correct order.
Job Templates
Just like with the fragments, the standard job files can be found here:
/chem/template/$NAME/
Most of the job files are just standard shell scripts with the headers required for the various clusters. The job files can be accessed by either the first letter of the cluster, or the full name (case insensitive).
Get User's Public Key
This is used mainly for getting the user's public key for use on the clusters. So instead of having to copy the file to the cluster and then append it to the authorized_keys file one can just wget and append.
/u/$USERNAME/id_rsa.pub
wget /u/$USERNAME/id_rsa.pub -O- >> ~/.ssh/authorized_keys
Get Running Jobs
This is used to allow viewing the currently running jobs of the logged in user. It returns two values. The first is is_authenticated which is used internally to determine whether or not the user is logged in. The second is clusters, an array of clusters objects that the user has added credentials for. Each one of those objects contains 3 properties: name, columns, jobs. name is the name of the cluster. columns is an array of all the column names. jobs is a 2D array of jobs from that cluster. The first dimension is all of the jobs themselves. The second is the properties. The properties are the same as what is given by the command qstat on the clusters split up based on spaces.
{
"is_authenticated": true,
"clusters": [
{
"jobs": [
["9969", "ccollins", "5b_TON_4c_6d.mjo", "59gb", "48:00:00", "--", "R"],
...
],
"name": "Marcy",
"columns": ["Job ID", "Username", "Jobname", "Req'd Memory", "Req'd Time", "Elap Time", "S"]
},
...
]
}